Tuesday, February 14, 2012
Traits vs. Genes
Hybrid Sunfish
Sunday, February 5, 2012
Software for large groups of IBD matches.
I received the following question, which I think is excellent.
Is there any software for plotting possible/probable relationships to MRCA [most recent common ancestor] for groups of 10-1,000 cousin matches suggested by various services, e.g. 23andMe, GEDmatch, FTDNA?
It seems to me that standard genealogical services are too rigid. Once you lay down a line of possible 3rd cousins and another for 4th cousins etc, linking them to various MRCAs who are perhaps 5 or 6 away becomes a bit of a nightmare.
The cousin matches referred to are typically based on shared segments longer than some threshold (typically 5 cM.) that appear to be identical by descent (IBD). In my case, 23andMe identifies 835 relatives but I know how I am related to only two (my mother and my sister).
I will contact scientists I know who work in this field to see if there is any software that is available to, and usable by, the general (informed, curious and sophisticated) public.
If you know of software that might be useful and want to see some of this data, I can send examples from 23andMe. It lists all IBD regions longer than the threshold that are shared with the user. Someone using this tool for genealogy will typically have such data for a small number of family members, and the information about most "cousins" is typically minimal. The person who posed this question can probably files to anyone who wants to see what they look like.
Saturday, February 12, 2011
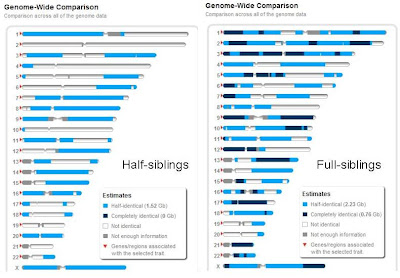
Half-siblings vs. Full-siblings visualized
Saturday, August 7, 2010
Avoiding overlaps in chromosome image (karyotypes)
I am doing a project in detecting the numerical abnormalities in chromosomes. Is it possible to get the microscopic chromosome images without any overlaps? ... mail me your suggestions.

I'm mailing you a sample image. The chromosomes in this image are overlapped and have crossovers. Is it possible to have the microscopic image without this type of crossovers. When I try to count the number of chromosomes the chromosomes that have crossovers and overlaps are counted as one. And my research concept is not mainly on this overlaps. So I'm trying for images without overlaps and crossovers. My concept is based on classification of the chromosomes so I need good well spread imageI have no training in cytogenetics per se, so I invite replies from those with such training (either academic human cytogeneticists or clinical laboratory specialists in cytogenetics (CLSp(CG)). As usual, answers should be provided in the form of comments from identified people with relevant expertise.
It seems to me that there are two questions here. One concerns the technical issue of whether it is possible to routinely obtain spreads without overlapping chromosomes. The other is whether there are already standard methods for dealing with this problem in the image analysis.
Sunday, July 4, 2010
Identifying half-siblings by genetic tests
1) Is it absolutely possible to tell if two females have the same father? These supposedly half sisters do not share the same mother. The father in question is deceased. Will our DNA reveal, certainly, that we share the same father? How do we find a reliable U.S. company to do this for us?
Information from a company like 23andme will provide you with enough markers to be completely sure. I'm not entirely sure how their interface works, so I can't tell you how easy the interpretation will be, but there will be enough data for you to be sure, and you will be able to get help with the interpretation. The reason is that two women who share a father will have an entire X chromosome in common. That is extremely unlikely to occur otherwise.
2) My sister donated a kidney to me. We were told that we matched 5 out of 6 genetic markers. From that information, is it possible to make an "educated guess" as to whether we are more likely to be full or half sisters? We believed we were half-sisters but some interesting coincidences lead me to believe that we may be full-sisters.
The six markers used for this test are not conclusive. As you know, even half-siblings can be a perfect match. A conclusive test would require many markers. Fortunately, companies like 23andme and Navigenetics provide information about many markers (about 450,000), and those tests could tell you definitively. If you are full siblings, then there will be parts of the genome (about one-fourth of the total) where you are identical for long stretches. That would be extremely unlikely if you are only half siblings. However, a small number of markers (less than 200 or so) would make it harder to make a definitive distinction between being half siblings and being full siblings. Six is definitely too few.
3) I wonder if you could help me. In trying to find my biological father, I came up with what could be two half siblings. The parents in both cases are deceased. I have been given a price of $500 for the three of us to test by saliva. Do you think without any parents, this could prove half siblings or would it be a waste of money?
You will share one of your two alleles with a half-sibling at about half of the sites in your genome. So, the answer is that with enough markers (thousands) the answer will be absolutely clear. The source of DNA (saliva, cheek swab, blood) does not matter much. 23andme will do about 450,000 markers for $400 (per person) and give you lots of additional information. The technology is pretty standard so other firms are probably OK. Just make sure that there are many markers (more than 100,000) and that you get access to the data (not just their interpretation of the data). Once you get your results you'll want to look for large regions of the genome where you and the putative half-sibling share markers. Of course, your putative half-siblings will have to agree to this analysis.
4) I heard that a recent study proved that men don't have half children but any children by the same man are full brothers and sisters irregardless of all different birth mothers. Is there a genetic truth to this?
Two children with the same father and different mothers are referred to as half-siblings. What you are referring to is almost certainly a legal or cultural distinction, not the sort of thing that can be proved by a study.
--------------------------------------
As usual, I invite additional answers in the form of comments. We are looking for answers from people with some expertise, and you will be asked to log in so that we know who you are (no anonymous answers).
Sunday, January 10, 2010
A beginner's guide to genetics
Unfortunately, I am not extremely familiar with books of this sort. I provide a link to the Amazon.com reviews of "A Beginner's Guide to Genetics and its Applications." Amazon recommends "Genetics for Dummies" and "Abraham Lincoln's DNA and Other Adventures in Genetics" as similar books. The latter sounds like an interesting read. I also try to recommend informative sites via my Gene Info web site.
I'm posting the question here in the hope that a reader will have suggestions. Although it would be nice to hear from non-experts, I will stick to the policy that while questions can be anonymous, comments (answers) are moderated, and I will only approves comments from people who identify themselves.